Introduction
Beta-lactam antibiotics are the common drugs in the treatment of bacterial infections (1). These antibiotics have been widely used since 1980 in order to treat serious infections caused by gram-negative bacteria but resistance against this group of antibiotics happened quickly around the world (2-4). Production of beta-lactamase enzymes is the main mechanism by which the resistance to these antibiotics occurs (5). These enzymes hydrolyze the core of beta-lactam antibiotics and finally inactivate them. The emergence of new antibiotics such as broad spectrum cephalosporins, aztreonams, and their wide usage in the treatment of bacterial infectious diseases has led to the presence of a new class of enzymes called extended-spectrum beta-lactamases (ESBL) (6, 7). ESBL enzymes are often coded by large plasmids that contain resistance genes to several antibiotics such as aminoglycosides, trimethoprim, sulfonamides, tetracycline, and chloramphenicol. The resistance to several antibiotics is an important feature of the Enterobacteriaceae strains producing ESBL (8).
ESBLs are classified into four main classes from A to D (Ambler classification). ESBL class A includes beta- lactamases such as CTX-M, TEM, SHV, and their subtypes. CTX-M enzymes constitute a specified category of beta-lactamases class A that is growing at a rapid rate. The first CTX-M enzyme that was isolated from clinical isolates was CTX-M-1 isolated from Enterobacterial strains in Europe in the late 1980s (9). After that, a lot of new variants of CTX-M were defined, so that more than 50 variants were described based on differences in the amino acid sequence that were divided into 5 groups or sub-types including CTX-M-1, CTX-M-2, CTX-M-8, CTX-M-9, and CTX-M-25 (10, 11). CTX-M beta-lactamases are part of natural ESBLs which hydrolyze cefotaxime or ceftriaxone more than ceftazidime. Therefore, most of the CTX-M-producing strains show significant resistance against ceftazidime and cefotaxime (12, 13).
The TEM-1 enzyme was originally found in E. coli isolated from blood culture of a Greece patient called Temoniera and hence was named TEM (13). This enzyme caused resistance to penicillin and first genera-tion cephalosporins such as cephalothin and cephalo-ridine (14). Various types of TEM beta-lactamases have been obtained through substituting amino acids in the activation site of TEM so that more than 130 types of TEM have been identified (15). The prevalence of some of these enzymes is different in various parts of the world (16). Today, the number of organisms producing TEM enzymes has been increased, while this has been considered as a crisis in the treatment of infections caused by these bacteria (17).
Since the prevalence of drug resistance is increasing and it has delayed the treatment of patients, especially those admitted to hospitals, the present research was performed to assess the detection and treatment of infections caused by E. coli strains being resistant to beta-lactam antibiotics in Tehran's Baghiyatallah Hospital.
Material and Methods
Isolation of Bacteria from Urine Specimens
This was a cross-sectional descriptive study. In a period of 6 months, 100 urine samples were collected from patients referred to Baghiyatallah Hospital of Tehran, and cultured on selective media of Eosin Methylene Blue (EMB) agar. Then, the plates were incubated at 37°C for 24 hours. The colonies grown underwent biochemical tests using MRVP broth, Simon citrate agar, TSI agar, SIM agar, MacConkey agar as well as Urea broth media. Finally, the E. coli isolates were identified and the corresponding colonies were kept at -70°C in the Skim Milk medium to be used in the later stages.
Identification of ESBL-producing E. coli Strains
The pattern of antibiotic susceptibility was studied by disk diffusion (Kriby-Bauer method) according to the instructions of Clinical and Laboratory Standards Institute (CLSI) using antibiotic disks manufactured by MAST Company (UK) including cefotaxime (30 mg), ceftriaxone (30 mg), ceftazidime (30 mg), cefpod-oxime (30 mg), nalidixic acid (10 mg), ceftizoxime (30 mg), cefixime (30 mg), nitrofurantoin (10 mg), ami-kacin (10 mg), and gentamicin (10 mg) (18). In order to perform disk diffusion, at first microbial suspensions were prepared as 0.5 McFarland from 18-hour colonies and then cultured on the surface of Mueller-Hinton agar (Merck, Germany), using a sterile swab. After 15 minutes, antibiotic disks were placed on the surface of the media every 20 mm from each other. After 15 min-utes, plates were incubated at 35 to 37°C for 16 to 18 hours. The diameter of inhibition zone was measured and interpreted according to the criteria of CLSI. Iso-lates with inhibition zone diameters of ≤22, ≤25, and ≤27 for ceftazidime, ceftriaxone, and cefotaxime, respectively, were evaluated for the presence of ESBL (18).
Phenotypic Confirmatory Test
To confirm ESBL production in candidate organisms, phenotypic confirmatory Double Disk Test (DDT) was used according to the CLSI command. The hybrid disks containing ceftazidime (30 mg) + clavulanic acid (10 mg) and cefotaxime (30 mg) + clavulanic acid (10 mg) were prepared from MAST Company. Afterward, the plates were incubated at 37°C for 24 hours. Then the diameter of inhibition zone was measured using a millimeter ruler and the results were interpreted according to the CLSI standards.
If the inhibition zone diameter of a colony around the combination disk was at least 5 mm bigger than the inhibition zone diameter of the single disk of the same antibiotic, it was considered as ESBL-producing isolate (18). In this test, the bacteria E. coli with the code of ATCC25922 and Klebsialla pneumonia with the code of ATCC700603 were used as ESBL positive and negative controls, respectively.
DNA Extraction and PCR
After performing the initial phenotypic con-firmatory test, the samples containing resistant isolates were selected for DNA extraction. Boiling method was used for DNA extraction (19) and PCR reaction was done to detect beta-lactamase genes including bla-CTX-M and bla-TEM under the conditions presented in Table 1. The forward and revers primers used in the present study were designed using the Primer3 software and their specificity was determined by Primer-blast online software (http://www.ncbi.gov/-tools/primer-blast). Moreover, to assure the quality of the primers, the Oligo analyzer program was used. The reaction mixture in a volume of 20 ml contained: 10 ml of master mix, 2 microliters of forward and revers primers (20, 21) (Table 2), 1 mL of extracted genome, and 7 mL of sterile distilled water.
Finally, electrophoresis of PCR products was performed in order to identify specific fragments with the sizes of 593bp (for blaCTX-M gene) and 867bp (for bla-TEM gene) on a 1.5% agarose gel with a size marker of 100 bp.
Results
The percentage of the identified isolates being resistant to the antibiotics are listed in Table 3.
Based on the results of disk diffusion screening test, 28 samples (28%) were resistant to cefotaxime, ceftazidime, and ceftriaxone, simultaneously. In DDT test, 21 cases were confirmed to be ESBL producing isolates.
According to the PCR results, the presence of blaCTX-M and blaTEM genes in 21 ESBL positive isolates were 21% and 20% for mentioned genes, respectively (Figures 1 and 2). Additionally, 19 samples (19%) contained isolates harboring both of the genes.
| Cycle repeats | Time (min) | Temperature (˚C) | Step | ||
|---|---|---|---|---|---|
| blaCTX-M | blaTEM | ||||
| 1 | 5 | 96 | 95 | 1. Initial denaturation | |
| 30 | 1 | 96 | 94 | Denaturation | 2. Amplification |
| 1 | 60 | 52.2 | Annealing | ||
| 1 | 72 | 72 | Extension | ||
| 1 | 7 | 72 | 72 | 3. Final Extension | |
| Primer | 5´-sequence-3´ | Expected-size (bp) | Reference |
|---|---|---|---|
| TEM-FTEM-R | ATGAGTATTCAACATTTCCGCTGACAGTTACCAATGCTTA | 867 | |
| CTX-M-FCTX-M-R | ATGTGCAGYACCAGTAARGTTGGGTRAARTARGTSACCAGA | 593 |
| Antibiotic | Number of resistant isolates (%) |
|---|---|
| Amoxicillin | 2 (16.7) |
| Amikacin | 4 (7.8) |
| Gentamicin | 12 (20.3) |
| Cephalexin | 5 (29.4) |
| Ceftazidime | 21 (35.0) |
| Ceftriaxone | 26 (41.3) |
| Cefotaxime | 4 (40.0) |
| Ceftizoxime | 2 (100) |
| Nitrofurantoin | 6 (9.7) |
| Nalidixic acid | 32 (64.0) |
| Ciprofloxacin | 22 (35.5) |
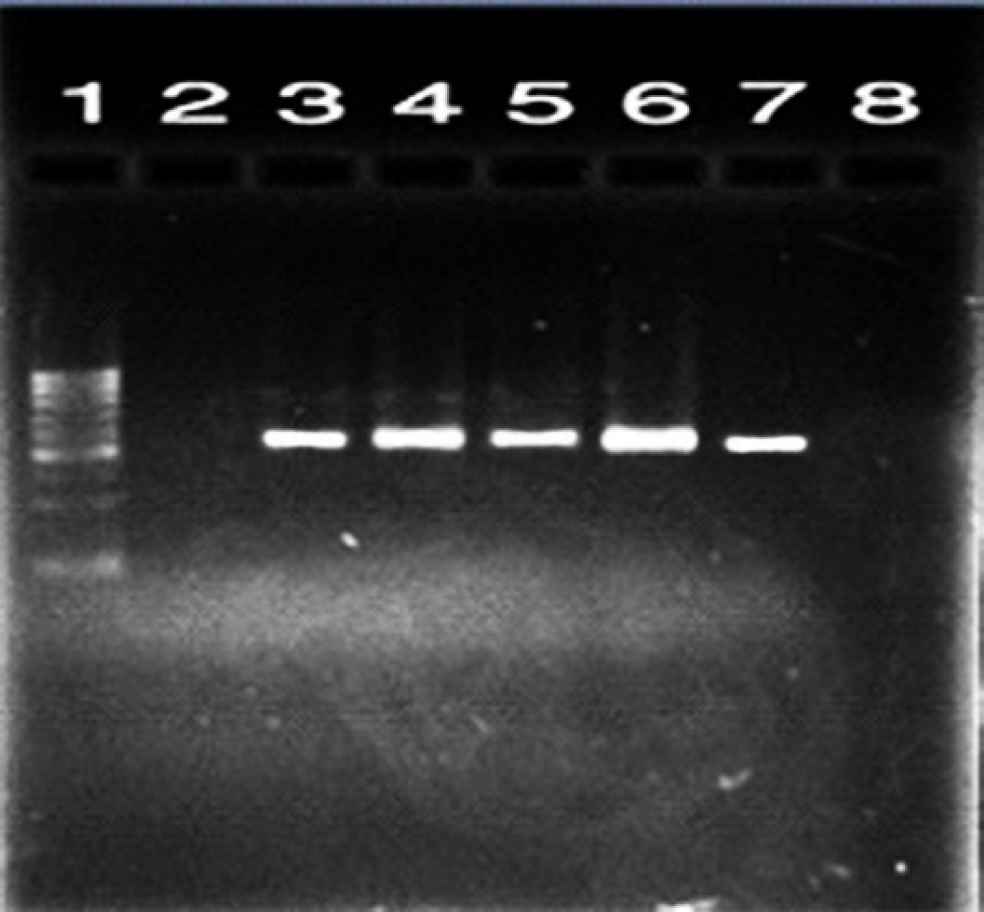
Fig. 1.Electrophoresis of PCR products for blaCTX-M gene. Lane 1: size marker (100 bp DNA ladder); lane 2: negative control, lane 3: positive control for blaCTX-M gene; lanes 4-7: PCR positive samples (593 bp) for blaCTX-M gene; lane 8: negative sample with no band
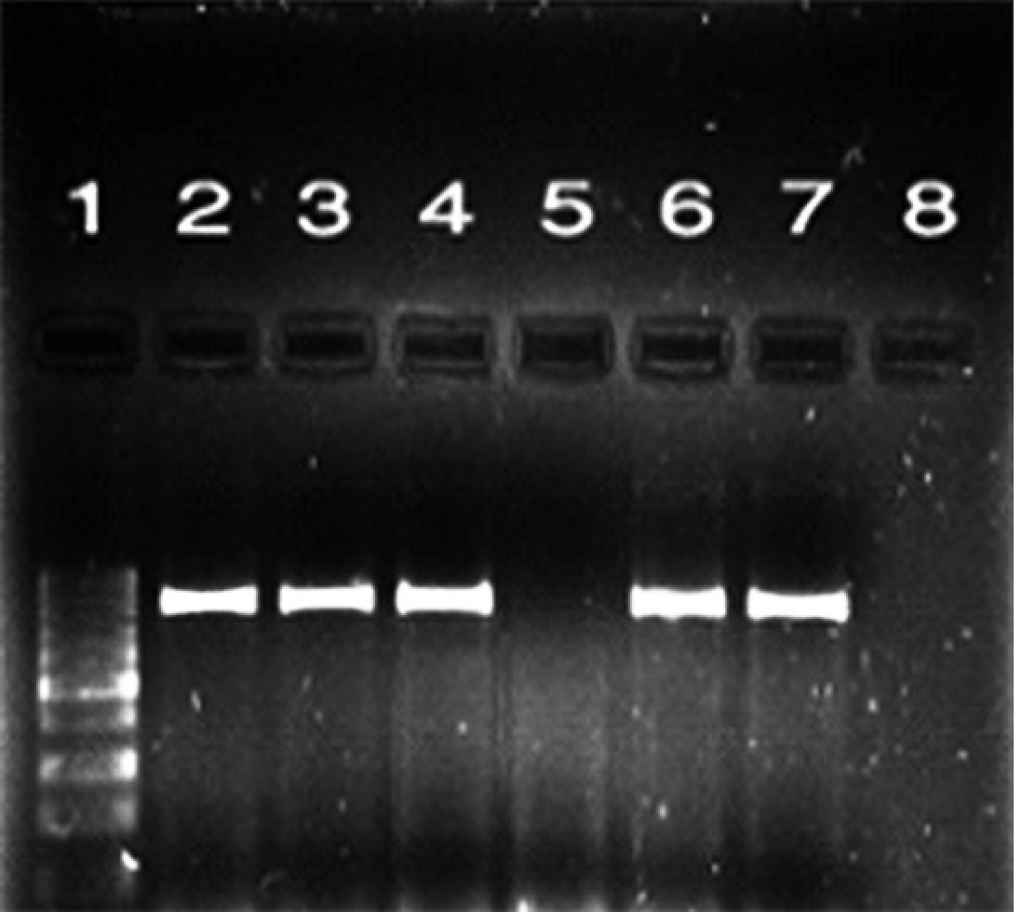
Fig. 2.Electrophoresis of PCR products for blaTEM gene. Lane 1: size marker (100 bp DNA ladder); lane 2: positive control for TEM gene; lanes 3, 4, 6, and 7: PCR positive samples for bla-TEM gene (867bp); lane 5: negative sample with no band; lane 8: negative control
Discussion
Bacterial beta-lactamase genes, especially ESBLs genes are among the effective factors that increase their resistance to beta-lactam antibiotics such as broad-spectrum cephalosporins. ESBL producing bacteria have created many health problems in recent years and novel methods are required to detect these bacteria in the clinical microbiology laboratories (21, 22). ESBL phenotypic detection is an appropriate method for differentiation between ESBL-producing isolates and isolates that use other mechanisms of beta-lactam antibiotics resistance (23). In the present study, among the analyzed samples, 21 samples (21%) were ESBL producers which indicate a high rate of ESBL-producing E. coli isolates in patients with urinary tract infection. These isolates were susceptible to the clavulanic acid (a broad spectrum beta-lactamase) and one case was not susceptible to this antibiotic which could be due to the production of enzymes such as AmpC (24). The prevalence of ESBL production in E. coli- and Klebsiella pneumonia-positive samples is different in various countries; for example, in Korea, the prevalence of these organisms is in the range of 4.8%-7.5% and 22.5%-22.8% for E. coli and Klebsiella pneumoniae, respectively. In India, the frequency of ESBL production is 34.2% and 27.3% for E.coli and Klebsiella pneumoniae, respectively (25). These results had no statistically significant difference compared to our results.
Several studies have indicated that ESBL production by nosocomial and non-nosocomial E. coli strains, has been quickly spread around the world, due to the emergence of ESBL type CTX_M. Lewis et al., in 2007 using phenotypic tests and molecular techniques, examined ESBL production on 94 urine samples of patients infected with uropathogenic E. coli and reported that among the samples, the most common ESBL type was CTX-M (26).
In the present research, in line with the other studies, susceptibility to gentamicin in the isolates producing CTX-M was more than that in the isolates producing TEM. Recent studies in Canada, Italy, Spain, Greece, and the United Kingdom revealed that ESBL production in E. coli, especially those that produce CTX-M have multiple resistances to trimethoprim, Sulfamethoxazole, tetracycline, gentamicin and ciprofloxacin (27).
Pitout et al. (2005) found that Enterobacteriaceae (like E. coli) isolated from urinary tract infections that produce ESBL type CTX-M are resistant to quinolones; in other words, resistance to quinolones is often associated with CTX-M (27). This relationship is confirmed in our study as in the isolates having CTX-M, 11 cases with resistance to ciprofloxacin were observed while was not observed in any other type of ESBLs. Moreover, the isolates in our study had the characteristics of ESBL-mediated resistance because a high percentage of isolates were resistant to the third generation cephalosporins, including: 35%, 41.3%, 40%, 100%, 50%, and 66.7% for ceftazidime, ceftriaxone, cefotaxime, ceftizoxime, cefixime and cefpodoxime, respectively. In a number of countries such as Iran, cephalosporins are the antibiotics of choice using to treat UTIs and practitioners use these antibiotics in abundance.
In a study conducted by Pour Akbari et al. in 2012, 100 E. coli strains isolated from UTI in patients aged 2 to 12 years underwent antibiotic susceptibility testing using disk diffusion method. Their results indicated a high percentage of susceptibility of E. coli to antibiotics including amikacin and nitrofurantoin (95% and 91%, respectively) (28). Meanwhile, in another research in the United States, the sensitivity to antibiotics obtained as 90% (29). We found similar results in our study regarding the susceptibility to the two antibiotics, amikacin (72.6%), and nitrofurantoin (83.9%).
Conclusion
The use of amikacin as empiric therapy is a good choice for patients with urinary tract infection as well as the use of nitrofurantoin as an antibiotic to prevent urinary tract infections. Moreover, concerning the production of ESBL by some isolates, phenotypic detection of ESBL-producing isolates is routinely suggested. The treatment should be selected regarding the ESBL production to avoid treatment failure.
Acknowledgements
None.
Conflict of Interest
The authors declared no conflicts of interest.
References
- Zaniani FR, Meshkat Z, Nasab MN, Khaje-Karamadini M, Ghazvini K, Rezaee A. The prevalence of TEM and SHV genes among extended-spectrum beta-lactamases producing Escherichia coli and Klebsiella pneumoniae. Iran J Basic Med Sci. 2012; 15(1):654-60.
- Bradford PA. Extended-spectrum β-lactamases in the 21st century: characterization, epidemiology, and detection of this important resistance threat. Clin Microbiol Rev. 2001; 14(4):933-51.
- Kim YK, Pai H, Lee HJ, Park SE, Choi EH, Kim J. Bloodstream infections by extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae in children: epidemiology and clinical outcome. Antimicrob Agents Chemother. 2002; 46(5):1481-91.
- Haj Ebrahim Tehrani F, Moradi M, Ghorbani N. Bacterial Etiology and Antibiotic Resistance Patterns in Neonatal Sepsis in Tehran during 2006-2014. Iran J Pathol. 2017; 12(4):356-61.
- Livermore DM. beta-Lactamases in laboratory and clinical resistance. Clin Microbiol Rev. 1995; 8(4):557-84.
- Tenover FC, Raney PM, Williams PP, Rasheed JK, Biddle JW, Oliver A. Evaluation of the NCCLS extended-spectrum beta-lactamase confirmation methods for Escherichia coli with isolates collected during Project ICARE. J Clin Microbiol. 2003; 41(7):3142-6.
- Darvishi M, Forootan M, Nazer MR, Karimi E, Noori M. Nosocomial Infections, Challenges and Threats: A Review Article. Iran J Med Microbiol. 2020; 14(2):162-81.
- Eliopoulos GM, Bush K. New β-lactamases in gram-negative bacteria: diversity and impact on the selection of antimicrobial therapy. Clin Infect Dis. 2001; 32(7):1085-9.
- Paterson DL, Bonomo RA. Extended-spectrum beta-lactamases: a clinical update. Clin Microbiol Rev. 2005; 18(4):657-86.
- Mirzaee M, Pourmand M, Chitsaz M, Mansouri S. Antibiotic resistance to third generation cephalosporins due to CTX-M-Type extended-spectrum β-lactamases in clinical isolates of Escherichia coli. Iran J Public Health. 2009; 38(1):10-7.
- Pallecchi L, Bartoloni A, Fiorelli C, Mantella A, Di Maggio T, Gamboa H. Rapid dissemination and diversity of CTX-M extended-spectrum beta-lactamase genes in commensal Escherichia coli isolates from healthy children from low-resource settings in Latin America. Antimicrob Agents Chemother. 2007; 51(8):2720-5.
- Chen Y, Delmas J, Sirot J, Shoichet B, Bonnet R. Atomic resolution structures of CTX-M beta-lactamases: extended spectrum activities from increased mobility and decreased stability. J Mol Biol. 2005; 348(2):349-62.
- Gupta V. An update on newer beta-lactamases. Indian J Med Res. 2007; 126(5):417-27.
- Mlynarczyk G, Mlynarczyk A, Bilewska A, Dukaczewska A, Golawski C, Kicman A. [High effectiveness of the method with cefpirome in detection of extended-spectrum beta-lactamases in different species of gram-negative bacilli]. Med Dosw Mikrobiol. 2006; 58(1):59-65.
- Zamanzad B, Deyham B, Nafisi M, Karimi A, Farokhi E. The Frequency of TEM-1 Gene in Extended Spectrum Beta Lactamases Producing Escherichia coli, Klebsiella pneumoniae and Enterobacter Strains Isolated from Hospital Clinical Samples Using PCR. Sci J Hamadan Univ Med Sci. 2008; 14(4):19-25.
- Perilli M, Dell'Amico E, Segatore B, de Massis MR, Bianchi C, Luzzaro F. Molecular characterization of extended-spectrum beta-lactamases produced by nosocomial isolates of Enterobacteriaceae from an Italian nationwide survey. J Clin Microbiol. 2002; 40(2):611-4.
- Haghi F, Zeighami H, Keramati N, Hemmati F, Hajiahmadi F. Frequency of TEM extended spectrum beta lactamase producing Escherichia coli in clinical specimens by phenotypic and molecular methods in Zanjan. Zanjan Univ Med Sci J. 2013; 21(85):55-63.
- CLSI. Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing; 21st Informational Supplement. M100-S21. CLSI Wayne. 2011.
- Queipo-Ortuño MI, Colmenero JDD, Macias M, Bravo MJ, Morata P. Preparation of bacterial DNA template by boiling and effect of immunoglobulin G as an inhibitor in real-time PCR for serum samples from patients with brucellosis. Clin Vaccine Immunol. 2008; 15(2):293-6.
- Oliver A, Weigel LM, Rasheed JK, McGowan JE Jr, Raney P, Tenover FC. Mechanisms of decreased susceptibility to cefpodoxime in Escherichia coli. Antimicrob Agents Chemother. 2002; 46(12):3829-36.
- Cavallo JD, Leblanc F, Fabre R, Fourticq-Esqueoute A. Group d'Etude le la Resistance de PaaB [Survey of the antibiotic sensitivity of Pseudomonas aeruginosa in France and the distribution of beta-lactam resistance mechanisms: the GERPB 1999 study]. Pathol Biol (Paris). 2001; 49(7):534-9.
- Rupp ME, Fey PD. Extended spectrum beta-lactamase (ESBL)-producing Enterobacteriaceae: considerations for diagnosis, prevention and drug treatment. Drugs. 2003; 63(4):353-65.
- Pitout JD, Hossain A, Hanson ND. Phenotypic and molecular detection of CTX-M-beta-lactamases produced by Escherichia coli and Klebsiella spp. J Clin Microbiol. 2004; 42(12):5715-21.
- Bell JM, Chitsaz M, Turnidge JD, Barton M, Walters LJ, Jones RN. Prevalence and significance of a negative extended-spectrum beta-lactamase (ESBL) confirmation test result after a positive ESBL screening test result for isolates of Escherichia coli and Klebsiella pneumoniae: results from the SENTRY Asia-Pacific Surveillance Program. J Clin Microbiol. 2007; 45(5):1478-82.
- Akram M, Shahid M, Khan AU. Etiology and antibiotic resistance patterns of community-acquired urinary tract infections in JNMC Hospital Aligarh, India. Ann Clin Microbiol Antimicrob. 2007; 6(1):4 .
- Lewis JS, Herrera M, Wickes B, Patterson JE, Jorgensen JH. First report of the emergence of CTX-M-type extended-spectrum β-lactamases (ESBLs) as the predominant ESBL isolated in a US health care system. Antimicrob Agents Chemother. 2007; 51(11):4015-21.
- Pitout JD, Nordmann P, Laupland KB, Poirel L. Emergence of Enterobacteriaceae producing extended-spectrum beta-lactamases (ESBLs) in the community. J Antimicrob Chemother. 2005; 56(1):52-9.
- Pourakbari B, Ferdosian F, Mahmoudi S, Teymuri M, Sabouni F, Heydari H. Increase resistant rates and ESBL production between E coli isolates causing urinary tract infection in young patients from Iran. Braz J Microbiol. 2012; 43(2):766-9.
- Mohammadi-Mehr M, Feizabadi M. Antimicrobial resistance pattern of Gram-negative bacilli isolated from patients at ICUs of Army hospitals in Iran. Iran J Microbiol. 2011; 3(1):26-30.

